Elizabeth Thrall

Assistant Professor
Physical/Biophysical Chemistry
Email: [email protected]
Office: JMH 632
Lab: JMH 628/630
Phone: 718-817-4495
-
- Assistant Professor at Fordham University
- NIH NRSA Postdoctoral Research Fellow at Harvard Medical School
- Ph.D. in Chemical Physics from Columbia University
- A.B. in Chemistry and Physics from Harvard University
-
1. Molecular Mechanisms of DNA Replication
My research uses single-molecule fluorescence microscopy, in combination with biochemical and microbiological approaches, to elucidate the molecular mechanisms of DNA replication in bacteria. Over the last few decades, researchers have constructed an increasingly detailed picture of DNA replication pathways in one bacterial species, the gram-negative model organism Escherichia coli. However, less is known about DNA replication in other bacteria. The goal of my research is to elucidate replication mechanisms in the model gram-positive species Bacillus subtilis, providing a more complete picture of bacterial DNA replication.
New single-molecule fluorescence imaging methods are uniquely suited for these studies. These imaging techniques use genetically-encoded fluorescent tags, including fluorescent proteins (FPs) (Figure 1A). Fusing a fluorescent tag to a protein of interest allows that protein to be visualized in the cell under laser excitation in a specialized microscope. One particularly powerful approach is single-particle tracking, such as particle-tracking photoactivation localization microscopy (sptPALM). This technique takes advantage of special photoactivatable fluorescent proteins (PAFPs), which are initially synthesized in a non-fluorescent (or dark) state but can be converted to a bright state by near-UV excitation. Using sptPALM, the localization, motion, and dynamics of single protein molecules can be measured in live cells (Figure 1B). As one example, the mobility of proteins in the cell can be quantified in the form of a diffusion coefficient (Figure 1C), providing information on the activity of a particular protein under conditions of interest.
Research in my lab is supported by the National Institute of General Medical Sciences of the National Institutes of Health through an R15 Academic Research Enhancement Award (AREA) (#R15GM151677). My research focuses on two main areas, but we also pursue collaborative projects related to DNA replication and repair:
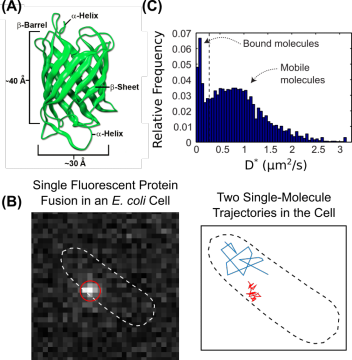
Figure 1. Single-molecule fluorescence imaging in live bacterial cells. (A) Structure of fluorescent proteins derived from Aequorea. Source: Shaner, N.C., et al. J. Cell. Sci. 2007, 120, 4227. (B) Fluorescence micrograph of a single fluorescent protein fusion in a live E. coli cell (left) and representative single-molecule trajectories in the same cell (right). (C) Distribution of apparent diffusion coefficients for the same protein in a live cell showing populations with distinct cellular mobilities.
A. The B. subtilis Replisome and Replicative Polymerase Coordination
From the simplest forms of life to the most complex, DNA replication is performed by a multi-protein machine called the replisome. Even in a relatively simple organism like E. coli, the replisome is composed of 13 different proteins, most of which are present in more than one copy (Figure 1A). Within the replisome, enzymes called DNA polymerases catalyze the synthesis of new DNA strands. Many bacteria, including E. coli, perform replication using a single replicative DNA polymerase, but other species require two polymerases. In B. subtilis, the polymerases PolC and DnaE are both essential for replication. In the current model, PolC performs the majority of DNA synthesis, but DnaE is needed to extend RNA primers before handing off the DNA template to PolC. This model requires frequent exchanges between PolC and DnaE, and it is unknown how the B. subtilis replisome coordinates the activity of these two polymerases. There is also evidence that DnaE may play a role in the response to DNA damage, although its function is unknown. My research is exploring the role of different protein-protein interactions in coordinating PolC and DnaE activity, both during normal replication and upon DNA damage.
B. Translesion Synthesis in B. subtilis
Although the replisome is highly efficient and accurate when copying undamaged DNA, it can be blocking by unrepaired DNA damage on the template strand, stalling replication and ultimately leading to cell death. As one way to resolve this type of stall, cells use a lower-fidelity replication pathway called translesion synthesis (TLS), in which specialized TLS polymerases copy damaged DNA (Figure 1B). Although TLS is important for cell survival upon DNA damage, these error-prone TLS polymerases can cause mutations; for that reason, their activity must be tightly regulated in the cell, especially in the absence of DNA damage. Studies of TLS in E. coli have revealed mechanisms by which its three TLS polymerases are regulated at multiple levels, but it is unknown whether these mechanisms are conserved across bacteria. My research is exploring the activity and regulation of the two TLS polymerases in B. subtilis, Pol Y1 and Pol Y2.
C. Other DNA Replication and Repair Pathways
In addition to my lab’s primary research areas, I am also engaged in collaborations on projects such as an investigation of DNA damage response pathways in E. coli and the development of peptide inhibitors for bacterial DNA replication.
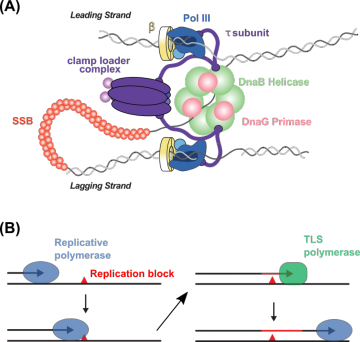
Figure 2. High and low-fidelity DNA replication. (A) Cartoon of the E. coli replisome. (B) Cartoon of high- and low-fidelity DNA polymerase exchange in translesion DNA synthesis.
2. Pedagogical Machine Learning and Artificial Intelligence Activities for Chemistry Students
Given the increasing ubiquity of computational approaches across STEM fields, I believe that it is critical to expose students to these methods early in their scientific careers. In particular, new approaches from artificial intelligence (AI), machine learning (ML), and data science (DS) have found numerous applications in chemistry in recent years, yet these topics are generally not included in chemistry curricula, and there is a paucity of pedagogical resources for instructors interested in including these topics in their courses. I am collaborating with chemistry and computer science faculty to develop new pedagogical activities to introduce students to AI/ML/DS in the context of chemistry. Although my initial projects have focused on activities for upper-level physical chemistry students, I am interested in extending this work to introductory chemistry courses in the future.
-
Please see Google Scholar for a complete list of publications.
O’Neal, Luke G.; Drucker, Madeline N.; Lai, Ngoc Khanh; Clemente, Ashley F.; Campbell, Alyssa P.; Way, Lindsey E.; Hong, Sinwoo; Holmes, Emily E.; Rancic, Sarah J.; Sawyer, Nicholas*; Wang, Xindan*; Thrall, Elizabeth S.* Nucleic Acids Res. 2025, doi: 10.1093/nar/gkaf721. “The B. subtilis Replicative Polymerases Bind the Sliding Clamp with Different Strengths to Tune Their Activity in DNA Replication.”
Marrin, McKayla E.; Foster, Michael R.; Santana, Chloe M.; Choi, Yoonhee; Jassal, Avtar S.; Rancic, Sarah J.; Greenwald, Carolyn R.; Drucker, Madeline N.; Feldman, Denholm T. Thrall, Elizabeth S.* Nucleic Acids Res. 2024, 52, 9613. “The Translesion Polymerase Pol Y1 is a Constitutive Component of the B. subtilis Replication Machinery.”
Thrall, Elizabeth S.*; Martinez Lopez, Fernando; Egg, Thomas J.; Lee, Seung Eun; Schrier, Joshua; Zhao, Yijun. J. Chem. Educ. 2023, 100, 4933. “Rediscovering the Particle-in-a-Box: Machine Learning Regression Analysis for Hypothesis Generation in Physical Chemistry Lab.”
Thrall, Elizabeth S.‡,*; Piatt, Sadie C.‡; Chang, Seungwoo; Loparo, Joseph J.* Proc. Natl. Acad. Sci. U.S.A. 2022, 119, e2208875119. “Replication Stalling Activates SSB for Recruitment of DNA Damage Tolerance Factors.”
Thrall, Elizabeth S.*; Lee, Seung Eun; Schrier, Joshua; Zhao, Yijun. J. Chem. Educ. 2021, 98, 3269. “Machine Learning for Functional Group Identification in Vibrational Spectroscopy: A Pedagogical Lab for Undergraduate Chemistry Students.
*: Corresponding author
‡: Equal contribution
Undergraduate and master’s students italicized